Logdet Method Profiling Across Matrix Sizes¶
This notebook profiles the runtime of different log-determinant strategies used in bayespecon.
The profiling matrices come from a regular polygon grid generated by the bayespecon.dgp module. Each size n is a grid side length producing an n × n rook-contiguity layout with n² spatial units; the spatial graph is built from the polygon GeoDataFrame returned by the DGP function.
Methods compared:
exact: symbolic determinant routeeigenvalue: spectral precompute routegrid_dense: dense-grid + spline interpolation (wasdense_grid/grid)grid_sparse: exact sparse-LU grid (legacy-style sparse-LU, wassparse_grid/full)sparse_spline: sparse-LU + spline interpolation (wasspline)trace_mc: Monte Carlo trace approximation (wasmc)grid_ilu: ILU-based approximate grid (wasilu/ichol)chebyshev: Chebyshev polynomial approximation via Clenshaw’s algorithm (Pace & LeSage 2004)
For each matrix size, we report:
setup time: build + compile callable logdet function
evaluation time: average cost to evaluate at many rho values
import time
from dataclasses import dataclass
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pytensor
import pytensor.tensor as pt
from libpysal import graph
from bayespecon import dgp
from bayespecon.logdet import make_logdet_fn
def make_grid_w(n_side: int) -> np.ndarray:
"""Create a row-standardized rook-contiguity matrix from an n_side x n_side polygon grid.
Uses ``dgp.simulate_sar`` with ``create_gdf=True`` to generate the polygon
geometry, then builds a contiguity graph from the returned GeoDataFrame.
"""
gdf = dgp.simulate_sar(n=n_side, create_gdf=True)
W = (
graph.Graph.build_contiguity(gdf, rook=True)
.transform("r")
.sparse.toarray()
.astype(np.float64)
)
return W
def compile_logdet_callable(W: np.ndarray, method: str, rho_min: float, rho_max: float):
"""Return a compiled callable f(rho) and its setup time in seconds."""
t0 = time.perf_counter()
rho = pt.scalar("rho")
expr = make_logdet_fn(W, method=method, rho_min=rho_min, rho_max=rho_max)(rho)
fn = pytensor.function([rho], expr)
setup_s = time.perf_counter() - t0
return fn, setup_s
def bench_eval_seconds(fn, rhos: np.ndarray, repeats: int = 5) -> float:
"""Median per-call evaluation latency in microseconds."""
run_times = []
for _ in range(repeats):
t0 = time.perf_counter()
for r in rhos:
_ = fn(float(r))
elapsed = time.perf_counter() - t0
run_times.append(elapsed / len(rhos))
return float(np.median(run_times))
@dataclass
class ProfileConfig:
# Grid side lengths; obs count = n_side². e.g. 6→36, 9→81, 11→121, 13→169, 16→256.
sizes: tuple[int, ...] = (10, 20, 25, 40, 50, 75)
methods: tuple[str, ...] = (
"exact",
"eigenvalue",
"grid_dense",
"grid_sparse",
"sparse_spline",
"trace_mc",
"grid_mc",
"grid_ilu",
"chebyshev",
)
method_max_n: dict = None
eval_points: int = 80
eval_repeats: int = 3
seed: int = 2026
def __post_init__(self):
if self.method_max_n is None:
self.method_max_n = {
"exact": 20,
"eigenvalue": 75,
"grid_dense": 40,
"grid_sparse": 75,
"sparse_spline": 75,
"trace_mc": 75,
"grid_ilu": 75,
"chebyshev": 75,
"grid_mc": 75,
}
cfg = ProfileConfig()
# spline/mc are defined on nonnegative rho ranges; others can use symmetric ranges.
method_rho_bounds = {
"exact": (-0.95, 0.95),
"eigenvalue": (-0.95, 0.95),
"grid_dense": (-0.95, 0.95),
"grid_sparse": (-0.95, 0.95),
"grid_ilu": (-0.95, 0.95),
"sparse_spline": (1e-5, 0.95),
"trace_mc": (1e-5, 0.95),
"chebyshev": (-0.95, 0.95),
"grid_mc": (1e-5, 0.95),
}
results = []
skipped = []
for n in cfg.sizes:
W = make_grid_w(n_side=n)
print(f"Profiling n_side={n} ({n * n} obs)...")
for method in cfg.methods:
if n > cfg.method_max_n[method]:
skipped.append(
{
"n_side": n,
"n_obs": n * n,
"method": method,
"reason": "above method_max_n cap",
}
)
continue
rho_min, rho_max = method_rho_bounds[method]
rho_grid = np.linspace(rho_min, rho_max, cfg.eval_points)
try:
fn, setup_s = compile_logdet_callable(
W, method, rho_min=rho_min, rho_max=rho_max
)
eval_s = bench_eval_seconds(fn, rho_grid, repeats=cfg.eval_repeats)
results.append(
{
"n_side": n,
"n_obs": n * n,
"method": method,
"rho_min": rho_min,
"rho_max": rho_max,
"setup_ms": 1e3 * setup_s,
"eval_us": 1e6 * eval_s,
}
)
except Exception as exc:
skipped.append(
{
"n_side": n,
"n_obs": n * n,
"method": method,
"reason": f"failed: {type(exc).__name__}: {exc}",
}
)
res = pd.DataFrame(results).sort_values(["method", "n_obs"]).reset_index(drop=True)
if not res.empty:
res["total_ms"] = res["setup_ms"] + (res["eval_us"] * cfg.eval_points / 1e3)
res
Profiling n_side=10 (100 obs)...
Profiling n_side=20 (400 obs)...
Profiling n_side=25 (625 obs)...
Profiling n_side=40 (1600 obs)...
Profiling n_side=50 (2500 obs)...
Profiling n_side=75 (5625 obs)...
| n_side | n_obs | method | rho_min | rho_max | setup_ms | eval_us | total_ms | |
|---|---|---|---|---|---|---|---|---|
| 0 | 10 | 100 | chebyshev | -0.95000 | 0.95 | 722.277719 | 3.978663 | 722.596012 |
| 1 | 20 | 400 | chebyshev | -0.95000 | 0.95 | 758.464173 | 4.359875 | 758.812963 |
| 2 | 25 | 625 | chebyshev | -0.95000 | 0.95 | 882.457647 | 4.382663 | 882.808260 |
| 3 | 40 | 1600 | chebyshev | -0.95000 | 0.95 | 1654.462255 | 4.404575 | 1654.814621 |
| 4 | 50 | 2500 | chebyshev | -0.95000 | 0.95 | 752.061951 | 3.941362 | 752.377260 |
| 5 | 75 | 5625 | chebyshev | -0.95000 | 0.95 | 911.477320 | 4.425988 | 911.831399 |
| 6 | 10 | 100 | eigenvalue | -0.95000 | 0.95 | 1285.116469 | 5.931537 | 1285.590992 |
| 7 | 20 | 400 | eigenvalue | -0.95000 | 0.95 | 54.599606 | 8.452613 | 55.275815 |
| 8 | 25 | 625 | eigenvalue | -0.95000 | 0.95 | 132.798277 | 9.585488 | 133.565116 |
| 9 | 40 | 1600 | eigenvalue | -0.95000 | 0.95 | 942.341594 | 16.344075 | 943.649120 |
| 10 | 50 | 2500 | eigenvalue | -0.95000 | 0.95 | 2963.920611 | 22.829400 | 2965.746963 |
| 11 | 75 | 5625 | eigenvalue | -0.95000 | 0.95 | 31491.998706 | 44.213250 | 31495.535766 |
| 12 | 10 | 100 | exact | -0.95000 | 0.95 | 2475.782028 | 89.436313 | 2482.936933 |
| 13 | 20 | 400 | exact | -0.95000 | 0.95 | 18.263063 | 1173.498425 | 112.142937 |
| 14 | 10 | 100 | grid_dense | -0.95000 | 0.95 | 2854.714416 | 6.282837 | 2855.217043 |
| 15 | 20 | 400 | grid_dense | -0.95000 | 0.95 | 81.349781 | 6.202550 | 81.845985 |
| 16 | 25 | 625 | grid_dense | -0.95000 | 0.95 | 160.974738 | 5.577775 | 161.420960 |
| 17 | 40 | 1600 | grid_dense | -0.95000 | 0.95 | 978.400555 | 5.942325 | 978.875941 |
| 18 | 10 | 100 | grid_ilu | -0.95000 | 0.95 | 111.006070 | 5.865175 | 111.475284 |
| 19 | 20 | 400 | grid_ilu | -0.95000 | 0.95 | 229.573024 | 5.998050 | 230.052868 |
| 20 | 25 | 625 | grid_ilu | -0.95000 | 0.95 | 326.078930 | 5.778525 | 326.541212 |
| 21 | 40 | 1600 | grid_ilu | -0.95000 | 0.95 | 736.189758 | 5.856925 | 736.658312 |
| 22 | 50 | 2500 | grid_ilu | -0.95000 | 0.95 | 1148.339431 | 6.054438 | 1148.823786 |
| 23 | 75 | 5625 | grid_ilu | -0.95000 | 0.95 | 2678.070275 | 5.898238 | 2678.542134 |
| 24 | 10 | 100 | grid_mc | 0.00001 | 0.95 | 611.702303 | 5.795175 | 612.165917 |
| 25 | 20 | 400 | grid_mc | 0.00001 | 0.95 | 44.763182 | 6.118275 | 45.252644 |
| 26 | 25 | 625 | grid_mc | 0.00001 | 0.95 | 48.480427 | 5.882963 | 48.951064 |
| 27 | 40 | 1600 | grid_mc | 0.00001 | 0.95 | 119.889783 | 6.190912 | 120.385056 |
| 28 | 50 | 2500 | grid_mc | 0.00001 | 0.95 | 83.681824 | 5.718150 | 84.139276 |
| 29 | 75 | 5625 | grid_mc | 0.00001 | 0.95 | 221.385197 | 5.852775 | 221.853419 |
| 30 | 10 | 100 | grid_sparse | -0.95000 | 0.95 | 675.971289 | 5.675325 | 676.425315 |
| 31 | 20 | 400 | grid_sparse | -0.95000 | 0.95 | 203.167481 | 6.371612 | 203.677210 |
| 32 | 25 | 625 | grid_sparse | -0.95000 | 0.95 | 289.113445 | 5.843887 | 289.580956 |
| 33 | 40 | 1600 | grid_sparse | -0.95000 | 0.95 | 647.071772 | 6.246637 | 647.571503 |
| 34 | 50 | 2500 | grid_sparse | -0.95000 | 0.95 | 1018.195648 | 5.982638 | 1018.674259 |
| 35 | 75 | 5625 | grid_sparse | -0.95000 | 0.95 | 2485.795270 | 5.912387 | 2486.268261 |
| 36 | 10 | 100 | sparse_spline | 0.00001 | 0.95 | 616.743570 | 6.021338 | 617.225277 |
| 37 | 20 | 400 | sparse_spline | 0.00001 | 0.95 | 54.185906 | 5.798175 | 54.649760 |
| 38 | 25 | 625 | sparse_spline | 0.00001 | 0.95 | 64.033661 | 5.759725 | 64.494439 |
| 39 | 40 | 1600 | sparse_spline | 0.00001 | 0.95 | 111.491204 | 5.829975 | 111.957602 |
| 40 | 50 | 2500 | sparse_spline | 0.00001 | 0.95 | 148.235111 | 5.763987 | 148.696230 |
| 41 | 75 | 5625 | sparse_spline | 0.00001 | 0.95 | 373.482329 | 5.708763 | 373.939030 |
| 42 | 10 | 100 | trace_mc | 0.00001 | 0.95 | 677.150311 | 4.427375 | 677.504501 |
| 43 | 20 | 400 | trace_mc | 0.00001 | 0.95 | 681.863229 | 4.332825 | 682.209855 |
| 44 | 25 | 625 | trace_mc | 0.00001 | 0.95 | 681.441494 | 3.930062 | 681.755899 |
| 45 | 40 | 1600 | trace_mc | 0.00001 | 0.95 | 717.659141 | 4.460062 | 718.015946 |
| 46 | 50 | 2500 | trace_mc | 0.00001 | 0.95 | 717.261441 | 3.869963 | 717.571038 |
| 47 | 75 | 5625 | trace_mc | 0.00001 | 0.95 | 855.243193 | 4.141588 | 855.574520 |
if res.empty:
raise RuntimeError("No profiling results were generated.")
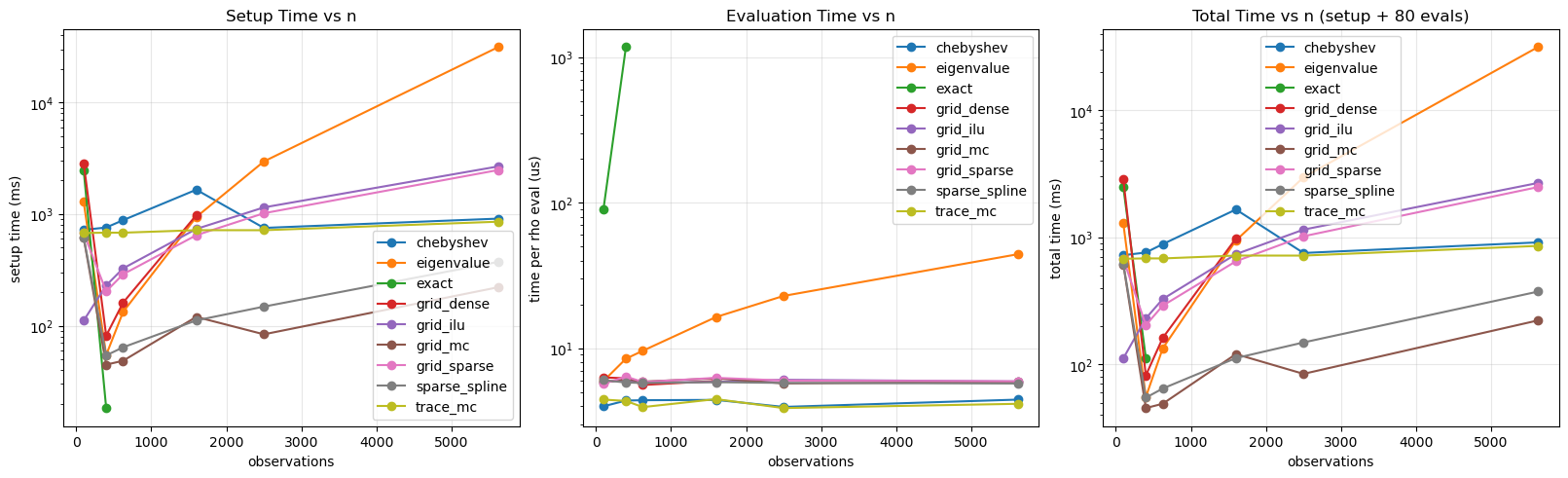
fig, axes = plt.subplots(1, 3, figsize=(16, 4.8), constrained_layout=True)
for method, grp in res.groupby("method"):
grp = grp.sort_values("n_obs")
axes[0].plot(grp["n_obs"], grp["setup_ms"], marker="o", label=method)
axes[1].plot(grp["n_obs"], grp["eval_us"], marker="o", label=method)
axes[2].plot(grp["n_obs"], grp["total_ms"], marker="o", label=method)
axes[0].set_title("Setup Time vs n")
axes[0].set_xlabel("observations")
axes[0].set_ylabel("setup time (ms)")
axes[0].set_yscale("log")
axes[0].grid(True, alpha=0.3)
axes[1].set_title("Evaluation Time vs n")
axes[1].set_xlabel("observations")
axes[1].set_ylabel("time per rho eval (us)")
axes[1].set_yscale("log")
axes[1].grid(True, alpha=0.3)
axes[2].set_title(f"Total Time vs n (setup + {cfg.eval_points} evals)")
axes[2].set_xlabel("observations")
axes[2].set_ylabel("total time (ms)")
axes[2].set_yscale("log")
axes[2].grid(True, alpha=0.3)
for ax in axes:
ax.legend()
plt.show()
summary = res.pivot_table(
index="n_obs", columns="method", values=["setup_ms", "eval_us", "total_ms"]
).sort_index()
display(summary)
if skipped:
skipped_df = (
pd.DataFrame(skipped).sort_values(["n_side", "method"]).reset_index(drop=True)
)
print("Skipped combinations (due to safety caps or failures):")
display(skipped_df)
| eval_us | setup_ms | total_ms | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| method | chebyshev | eigenvalue | exact | grid_dense | grid_ilu | grid_mc | grid_sparse | sparse_spline | trace_mc | chebyshev | ... | trace_mc | chebyshev | eigenvalue | exact | grid_dense | grid_ilu | grid_mc | grid_sparse | sparse_spline | trace_mc |
| n_obs | |||||||||||||||||||||
| 100 | 3.978663 | 5.931537 | 89.436313 | 6.282837 | 5.865175 | 5.795175 | 5.675325 | 6.021338 | 4.427375 | 722.277719 | ... | 677.150311 | 722.596012 | 1285.590992 | 2482.936933 | 2855.217043 | 111.475284 | 612.165917 | 676.425315 | 617.225277 | 677.504501 |
| 400 | 4.359875 | 8.452613 | 1173.498425 | 6.202550 | 5.998050 | 6.118275 | 6.371612 | 5.798175 | 4.332825 | 758.464173 | ... | 681.863229 | 758.812963 | 55.275815 | 112.142937 | 81.845985 | 230.052868 | 45.252644 | 203.677210 | 54.649760 | 682.209855 |
| 625 | 4.382663 | 9.585488 | NaN | 5.577775 | 5.778525 | 5.882963 | 5.843887 | 5.759725 | 3.930062 | 882.457647 | ... | 681.441494 | 882.808260 | 133.565116 | NaN | 161.420960 | 326.541212 | 48.951064 | 289.580956 | 64.494439 | 681.755899 |
| 1600 | 4.404575 | 16.344075 | NaN | 5.942325 | 5.856925 | 6.190912 | 6.246637 | 5.829975 | 4.460062 | 1654.462255 | ... | 717.659141 | 1654.814621 | 943.649120 | NaN | 978.875941 | 736.658312 | 120.385056 | 647.571503 | 111.957602 | 718.015946 |
| 2500 | 3.941362 | 22.829400 | NaN | NaN | 6.054438 | 5.718150 | 5.982638 | 5.763987 | 3.869963 | 752.061951 | ... | 717.261441 | 752.377260 | 2965.746963 | NaN | NaN | 1148.823786 | 84.139276 | 1018.674259 | 148.696230 | 717.571038 |
| 5625 | 4.425988 | 44.213250 | NaN | NaN | 5.898238 | 5.852775 | 5.912387 | 5.708763 | 4.141588 | 911.477320 | ... | 855.243193 | 911.831399 | 31495.535766 | NaN | NaN | 2678.542134 | 221.853419 | 2486.268261 | 373.939030 | 855.574520 |
6 rows × 27 columns
Skipped combinations (due to safety caps or failures):
| n_side | n_obs | method | reason | |
|---|---|---|---|---|
| 0 | 25 | 625 | exact | above method_max_n cap |
| 1 | 40 | 1600 | exact | above method_max_n cap |
| 2 | 50 | 2500 | exact | above method_max_n cap |
| 3 | 50 | 2500 | grid_dense | above method_max_n cap |
| 4 | 75 | 5625 | exact | above method_max_n cap |
| 5 | 75 | 5625 | grid_dense | above method_max_n cap |
Coefficient and Fit-Time Comparison Across Logdet Methods¶
This section uses a regular polygon grid generated by bayespecon.dgp to simulate one SAR dataset, maps the simulated response, and estimates the same SAR model using each logdet_method.
We compare:
posterior mean coefficients (
rho,beta_0,beta_1,beta_2)total wall-clock time to estimate each model
To keep this section runnable in docs contexts, sampling is intentionally modest.
from bayespecon import SAR
def simulate_sar_data(n_side: int = 16, seed: int = 2026):
"""Simulate SAR data on an n_side x n_side polygon grid using the DGP module."""
rng = np.random.default_rng(seed)
beta_true = np.array([1.0, 0.8, -0.5], dtype=np.float64)
rho_true = 0.35
sigma_true = 0.7
gdf = dgp.simulate_sar(
n=n_side,
rho=rho_true,
beta=beta_true,
sigma=sigma_true,
rng=rng,
create_gdf=True,
)
# Keep the SAR parameterization consistent with model assumptions.
W_graph = graph.Graph.build_contiguity(gdf, rook=True).transform("r")
y = gdf["y"].to_numpy()
X_cols = [c for c in gdf.columns if c.startswith("X_")]
X = gdf[X_cols].to_numpy()
return gdf, y, X, W_graph, rho_true, beta_true
def fit_sar_for_method(
y,
X,
W,
method: str,
draws: int = 400,
tune: int = 400,
seed: int = 2026,
rho_lower: float = 0.000001,
rho_upper: float = 0.95,
):
"""Fit SAR with a specific logdet method and return posterior means + runtime."""
t0 = time.perf_counter()
model = SAR(
y=y,
X=X,
W=W,
logdet_method=method,
priors={"rho_lower": rho_lower, "rho_upper": rho_upper},
)
idata = model.fit(
draws=draws,
tune=tune,
chains=2,
cores=1,
random_seed=seed,
target_accept=0.95,
progressbar=False,
compute_convergence_checks=False,
)
elapsed_s = time.perf_counter() - t0
beta_mean = idata.posterior["beta"].mean(("chain", "draw")).to_numpy()
rho_mean = float(idata.posterior["rho"].mean(("chain", "draw")).to_numpy())
return {
"method": method,
"total_time_s": elapsed_s,
"rho": rho_mean,
"beta_0": float(beta_mean[0]),
"beta_1": float(beta_mean[1]),
"beta_2": float(beta_mean[2]),
}
methods_for_model = [
"eigenvalue",
"sparse_spline",
"trace_mc",
"grid_mc",
"grid_ilu",
"chebyshev",
]
gdf_model, y_model, X_model, W_model, rho_true, beta_true = simulate_sar_data(n_side=25)
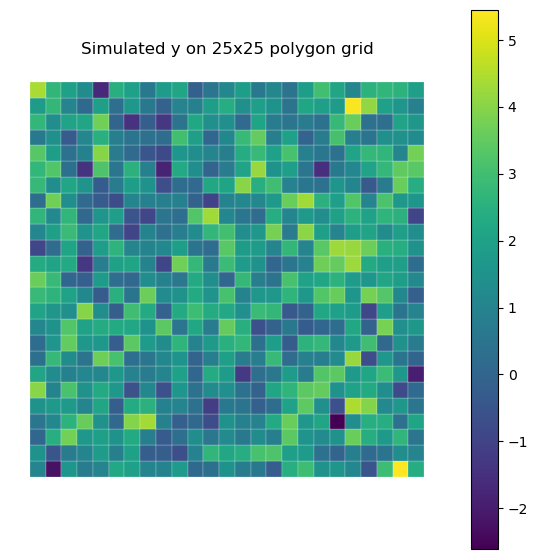
fig, ax = plt.subplots(1, 1, figsize=(7, 7))
gdf_model.plot(
column="y", cmap="viridis", legend=True, linewidth=0.15, edgecolor="white", ax=ax
)
ax.set_title("Simulated y on 25x25 polygon grid")
ax.set_axis_off()
plt.show()
model_rows = []
for m in methods_for_model:
print(f"Estimating SAR with method={m}...")
try:
model_rows.append(fit_sar_for_method(y_model, X_model, W_model, method=m))
except Exception as exc:
model_rows.append(
{
"method": m,
"total_time_s": np.nan,
"rho": np.nan,
"beta_0": np.nan,
"beta_1": np.nan,
"beta_2": np.nan,
"error": f"{type(exc).__name__}: {exc}",
}
)
coef_compare = pd.DataFrame(model_rows)
coef_compare = coef_compare.sort_values("total_time_s", na_position="last").reset_index(
drop=True
)
coef_compare["rho_true"] = rho_true
coef_compare["abs_err_rho"] = (coef_compare["rho"] - rho_true).abs()
for j, btrue in enumerate(beta_true):
coef_compare[f"beta_{j}_true"] = btrue
coef_compare[f"abs_err_beta_{j}"] = (coef_compare[f"beta_{j}"] - btrue).abs()
if "eigenvalue" in coef_compare["method"].values:
base = coef_compare.loc[
coef_compare["method"] == "eigenvalue", ["rho", "beta_0", "beta_1", "beta_2"]
].iloc[0]
for col in ["rho", "beta_0", "beta_1", "beta_2"]:
coef_compare[f"delta_vs_eigen_{col}"] = coef_compare[col] - base[col]
display(coef_compare)
Estimating SAR with method=eigenvalue...
Estimating SAR with method=sparse_spline...
Estimating SAR with method=trace_mc...
Estimating SAR with method=grid_mc...
Estimating SAR with method=grid_ilu...
Estimating SAR with method=chebyshev...
Initializing NUTS using jitter+adapt_diag...
Sequential sampling (2 chains in 1 job)
NUTS: [rho, beta, sigma]
Sampling 2 chains for 400 tune and 400 draw iterations (800 + 800 draws total) took 1 seconds.
Initializing NUTS using jitter+adapt_diag...
Sequential sampling (2 chains in 1 job)
NUTS: [rho, beta, sigma]
Sampling 2 chains for 400 tune and 400 draw iterations (800 + 800 draws total) took 1 seconds.
Initializing NUTS using jitter+adapt_diag...
Sequential sampling (2 chains in 1 job)
NUTS: [rho, beta, sigma]
Sampling 2 chains for 400 tune and 400 draw iterations (800 + 800 draws total) took 1 seconds.
Initializing NUTS using jitter+adapt_diag...
Sequential sampling (2 chains in 1 job)
NUTS: [rho, beta, sigma]
Sampling 2 chains for 400 tune and 400 draw iterations (800 + 800 draws total) took 1 seconds.
Initializing NUTS using jitter+adapt_diag...
Sequential sampling (2 chains in 1 job)
NUTS: [rho, beta, sigma]
Sampling 2 chains for 400 tune and 400 draw iterations (800 + 800 draws total) took 1 seconds.
Initializing NUTS using jitter+adapt_diag...
Sequential sampling (2 chains in 1 job)
NUTS: [rho, beta, sigma]
Sampling 2 chains for 400 tune and 400 draw iterations (800 + 800 draws total) took 1 seconds.
| method | total_time_s | rho | beta_0 | beta_1 | beta_2 | rho_true | abs_err_rho | beta_0_true | abs_err_beta_0 | beta_1_true | abs_err_beta_1 | beta_2_true | abs_err_beta_2 | delta_vs_eigen_rho | delta_vs_eigen_beta_0 | delta_vs_eigen_beta_1 | delta_vs_eigen_beta_2 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | grid_mc | 3.374331 | 0.381977 | 0.882365 | 0.809504 | -0.519954 | 0.35 | 0.031977 | 1.0 | 0.117635 | 0.8 | 0.009504 | -0.5 | 0.019954 | 5.195275e-04 | -2.154732e-03 | 9.726491e-04 | -1.701039e-03 |
| 1 | grid_ilu | 3.489972 | 0.382076 | 0.882487 | 0.807844 | -0.520888 | 0.35 | 0.032076 | 1.0 | 0.117513 | 0.8 | 0.007844 | -0.5 | 0.020888 | 6.188300e-04 | -2.032753e-03 | -6.876095e-04 | -2.634321e-03 |
| 2 | sparse_spline | 4.127959 | 0.382676 | 0.882155 | 0.808407 | -0.518637 | 0.35 | 0.032676 | 1.0 | 0.117845 | 0.8 | 0.008407 | -0.5 | 0.018637 | 1.219458e-03 | -2.364459e-03 | -1.241932e-04 | -3.838977e-04 |
| 3 | eigenvalue | 4.510765 | 0.381457 | 0.884519 | 0.808531 | -0.518253 | 0.35 | 0.031457 | 1.0 | 0.115481 | 0.8 | 0.008531 | -0.5 | 0.018253 | 0.000000e+00 | 0.000000e+00 | 0.000000e+00 | 0.000000e+00 |
| 4 | chebyshev | 8.506022 | 0.381457 | 0.884520 | 0.808532 | -0.518253 | 0.35 | 0.031457 | 1.0 | 0.115480 | 0.8 | 0.008532 | -0.5 | 0.018253 | -2.763719e-07 | 4.041846e-07 | 2.492285e-08 | -1.546428e-08 |
| 5 | trace_mc | 10.553827 | 0.381492 | 0.884468 | 0.808528 | -0.518253 | 0.35 | 0.031492 | 1.0 | 0.115532 | 0.8 | 0.008528 | -0.5 | 0.018253 | 3.492415e-05 | -5.178650e-05 | -3.062972e-06 | 8.995280e-07 |
Notes¶
exactis expected to scale poorly and is intentionally capped at smaller n in this notebook.eigenvalueusually has moderate setup cost and very fast repeated evaluations.grid_densehas an upfront grid-construction cost but can be competitive for repeated evaluation scenarios (wasdense_grid/grid).grid_sparseandsparse_splineare sparse-LU variants inspired by the legacy toolbox routines (werefullandint).trace_mcis stochastic and may vary slightly run-to-run unless random seeds are fixed.grid_iluuses an ILU approximation that can trade precision for speed depending on sparsity (wasilu/ichol).chebyshevuses Chebyshev polynomial approximation via Clenshaw’s algorithm (Pace & LeSage 2004). It pre-computes coefficients at Chebyshev nodes (O(n³) for eigenvalue path, O(R·n·m) for MC path) and then evaluates in O(m) per call. It supports negative rho ranges, unlikesparse_splineandtrace_mc.Parameter ranges:
sparse_splineandtrace_mcare profiled on a nonnegative range (rho in [1e-5, 0.95]), whileexact,eigenvalue,grid_dense,grid_sparse,grid_ilu, andchebyshevare profiled on a symmetric range (rho in [-0.95, 0.95]).Adjust
ProfileConfig.sizesandmethod_max_nfor deeper stress tests.
import arviz as az
import pandas as pd
# Assuming you have a list of inference data objects (idata) for each method,
# or you can refit/collect them as needed. If you only have the summary DataFrames,
# adapt the code to use those directly.
ess_rows = []
for i, m in enumerate(methods_for_model):
model = SAR(
y=y_model,
X=X_model,
W=W_model,
logdet_method=m,
priors={"rho_lower": 1e-5, "rho_upper": 0.95},
)
idata = model.fit(
draws=400,
tune=400,
chains=2,
cores=1,
random_seed=2026,
target_accept=0.95,
progressbar=False,
compute_convergence_checks=False,
)
summary = az.summary(idata, var_names=["rho"])
ess = summary.loc["rho", "ess_bulk"]
ess_rows.append({"method": m, "ess_rho": ess})
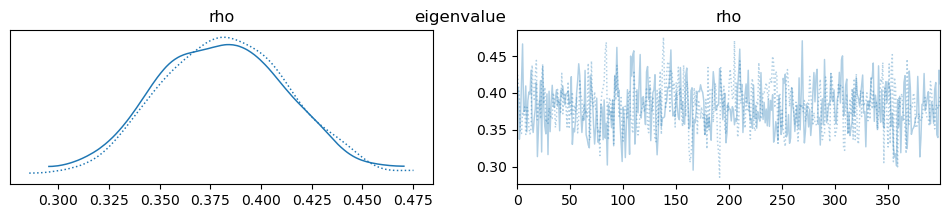
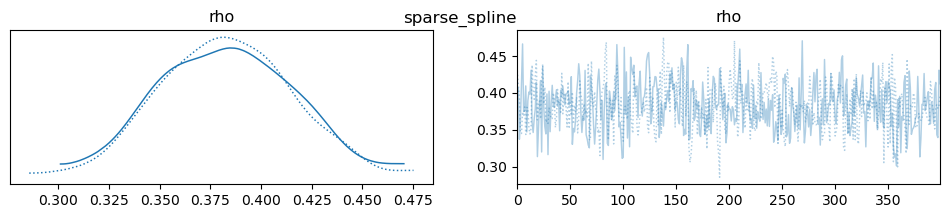
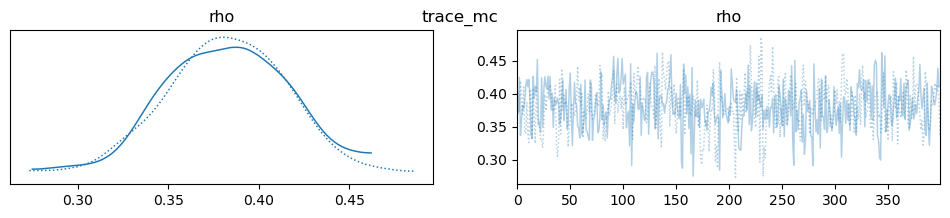
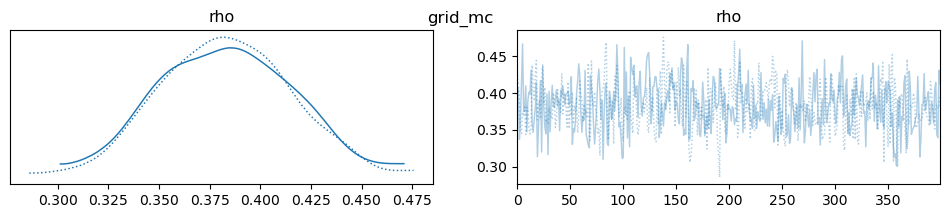
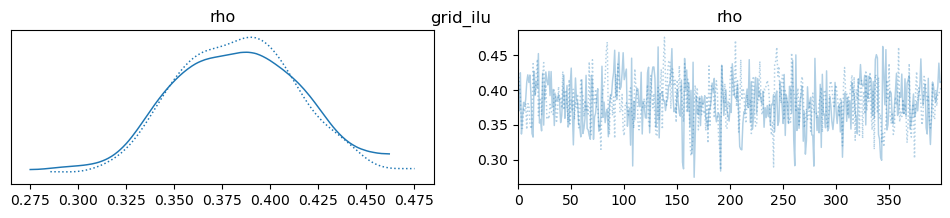
az.plot_trace(idata, var_names=["rho"], compact=True, legend=False)
plt.suptitle(m)
ess_df = pd.DataFrame(ess_rows)
display(ess_df)
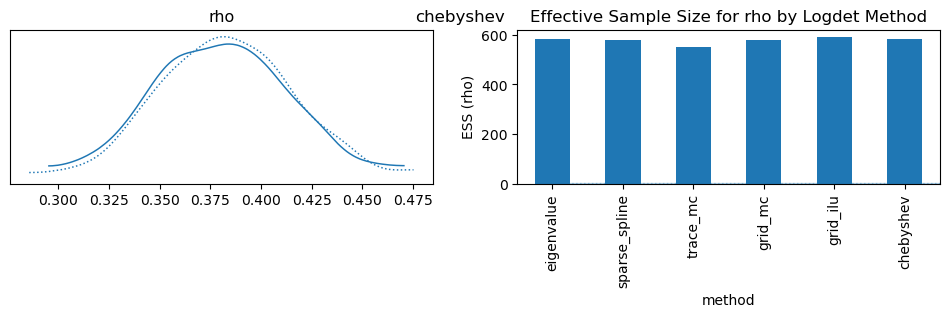
ess_df.set_index("method")["ess_rho"].plot.bar(
ylabel="ESS (rho)", title="Effective Sample Size for rho by Logdet Method"
)
Initializing NUTS using jitter+adapt_diag...
Sequential sampling (2 chains in 1 job)
NUTS: [rho, beta, sigma]
Sampling 2 chains for 400 tune and 400 draw iterations (800 + 800 draws total) took 1 seconds.
Initializing NUTS using jitter+adapt_diag...
Sequential sampling (2 chains in 1 job)
NUTS: [rho, beta, sigma]
Sampling 2 chains for 400 tune and 400 draw iterations (800 + 800 draws total) took 1 seconds.
Initializing NUTS using jitter+adapt_diag...
Sequential sampling (2 chains in 1 job)
NUTS: [rho, beta, sigma]
Sampling 2 chains for 400 tune and 400 draw iterations (800 + 800 draws total) took 1 seconds.
Initializing NUTS using jitter+adapt_diag...
Sequential sampling (2 chains in 1 job)
NUTS: [rho, beta, sigma]
Sampling 2 chains for 400 tune and 400 draw iterations (800 + 800 draws total) took 1 seconds.
Initializing NUTS using jitter+adapt_diag...
Sequential sampling (2 chains in 1 job)
NUTS: [rho, beta, sigma]
Sampling 2 chains for 400 tune and 400 draw iterations (800 + 800 draws total) took 1 seconds.
Initializing NUTS using jitter+adapt_diag...
Sequential sampling (2 chains in 1 job)
NUTS: [rho, beta, sigma]
Sampling 2 chains for 400 tune and 400 draw iterations (800 + 800 draws total) took 1 seconds.
| method | ess_rho | |
|---|---|---|
| 0 | eigenvalue | 585.0 |
| 1 | sparse_spline | 580.0 |
| 2 | trace_mc | 551.0 |
| 3 | grid_mc | 579.0 |
| 4 | grid_ilu | 591.0 |
| 5 | chebyshev | 585.0 |
<Axes: title={'center': 'Effective Sample Size for rho by Logdet Method'}, xlabel='method', ylabel='ESS (rho)'>
Method selection policy¶
Default for most work
Use chebyshev.
Reason: good speed, stable behavior, and supports both negative and positive rho ranges.
2.Exploratory runs where speed is most important
If rho is constrained to nonnegative values, use mc.
Keep this for quick iteration, model search, and early diagnostics.
Deterministic interpolation alternative
If rho is nonnegative and you want deterministic behavior, use spline.
Only use when prior bounds are fully inside the interpolation interval.
Final reporting and publication runs
Prefer chebyshev.
If using mc or spline, validate against chebyshev on the same data and priors.
Guardrails
If prior includes negative rho, do not use mc or spline.
Avoid switching methods mid-comparison unless you explicitly document it.
Record method choice and rho bounds in every benchmark or model report.
Suggested defaults by stage
Development: mc (nonnegative rho only), otherwise chebyshev. Pre-final validation: chebyshev. Final results: chebyshev, or mc/spline with chebyshev cross-check.